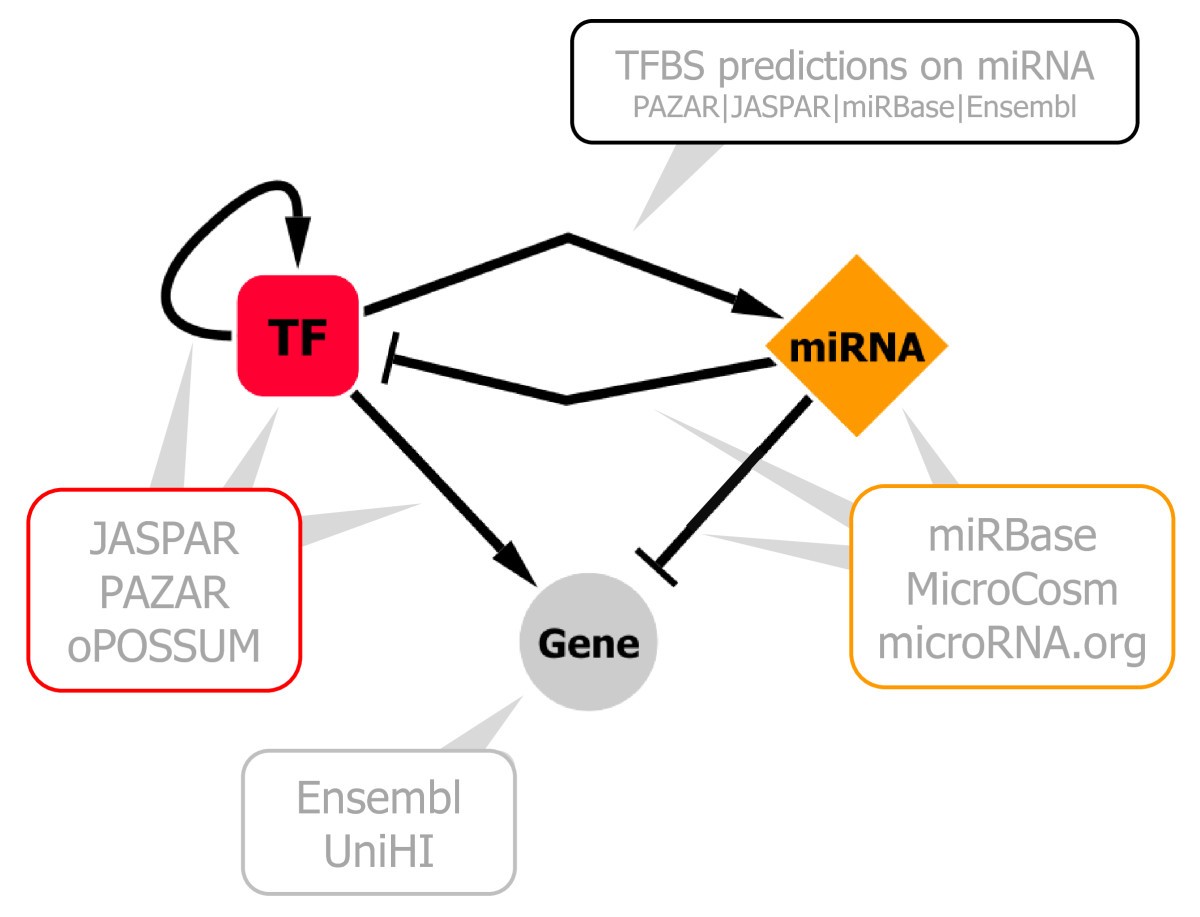
MIR@NT@N: a framework integrating transcription factors, microRNAs and their targets to identify sub-network motifs in a meta-regulation network model | BMC Bioinformatics | Full Text
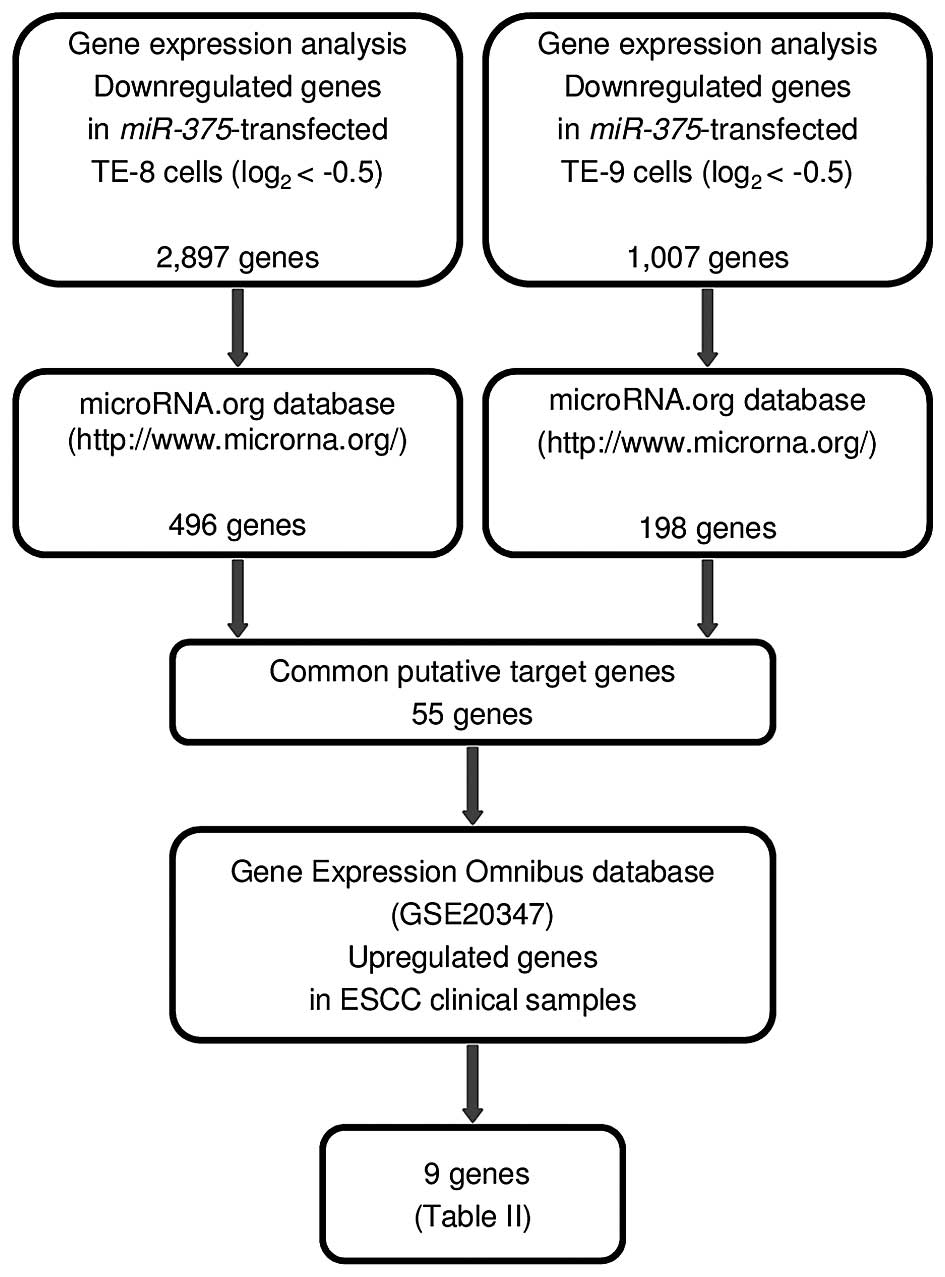
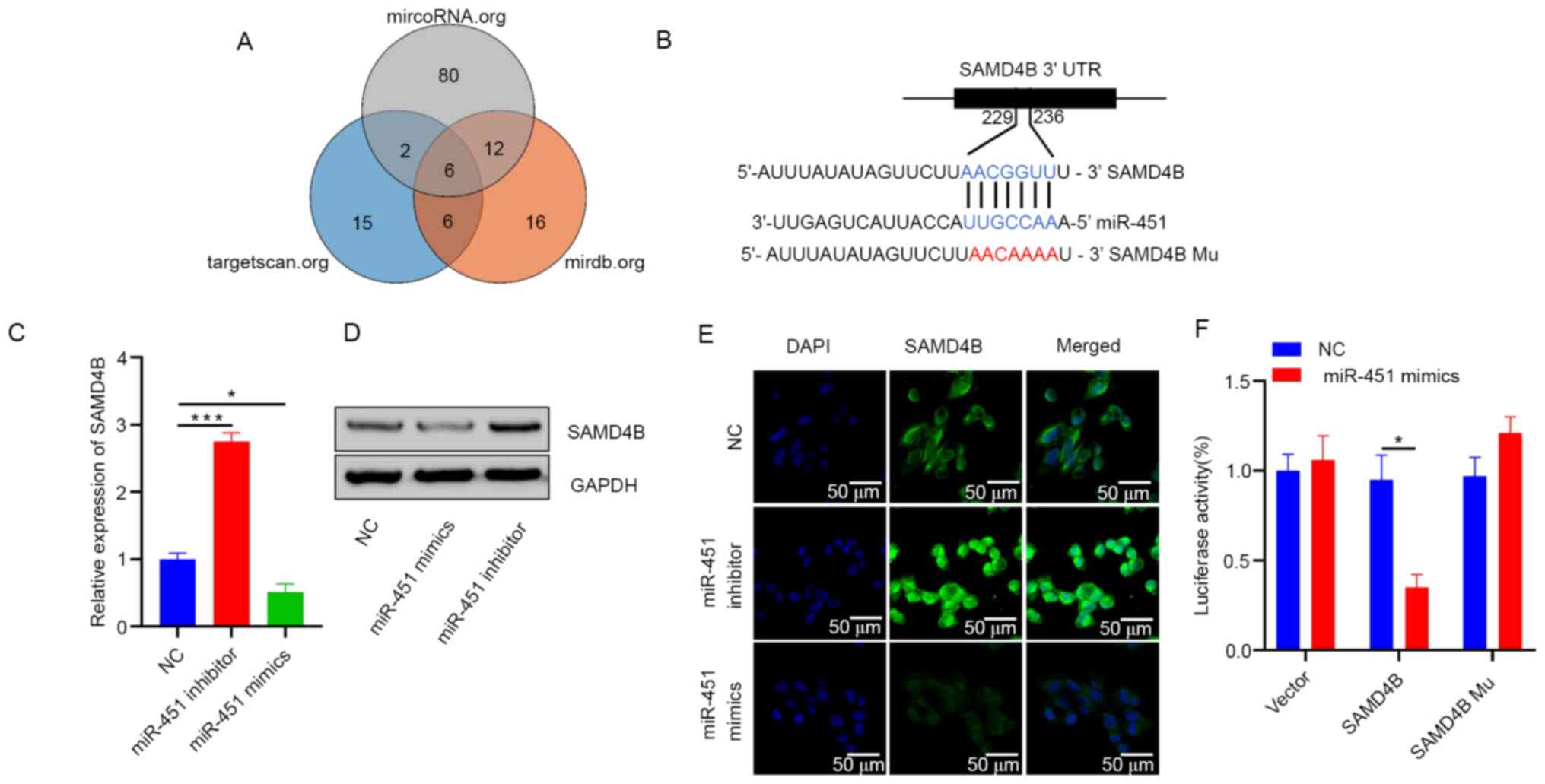
Regulation of MMP13 by antitumor microRNA-375 markedly inhibits cancer cell migration and invasion in esophageal squamous cell carcinoma

Manganese-dependent microRNA trimming by 3′→5′ exonucleases generates 14-nucleotide or shorter tiny RNAs | PNAS
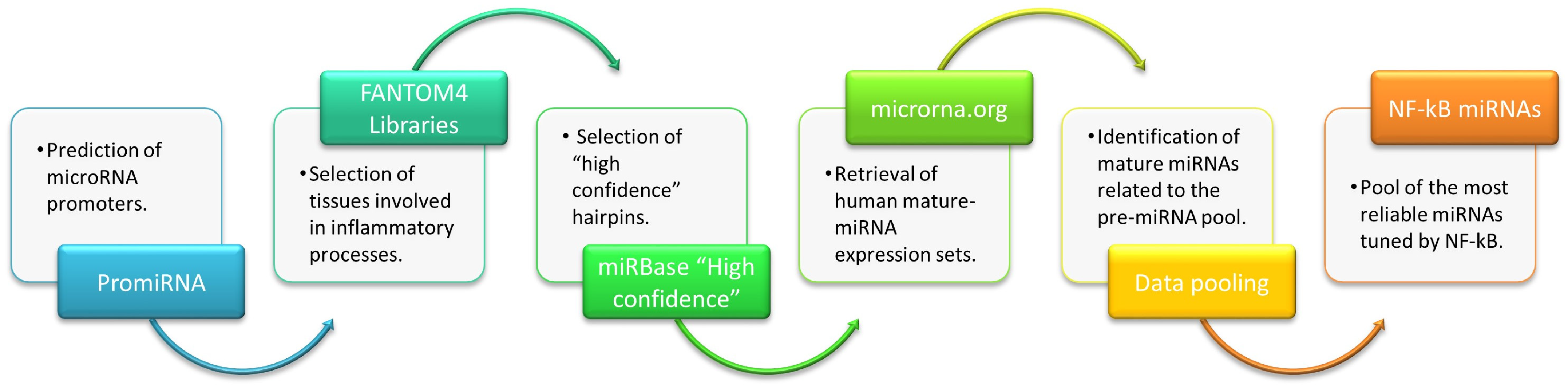
IJMS | Free Full-Text | A Data-Mining Approach to Identify NF-kB-Responsive microRNAs in Tissues Involved in Inflammatory Processes: Potential Relevance in Age-Related Diseases
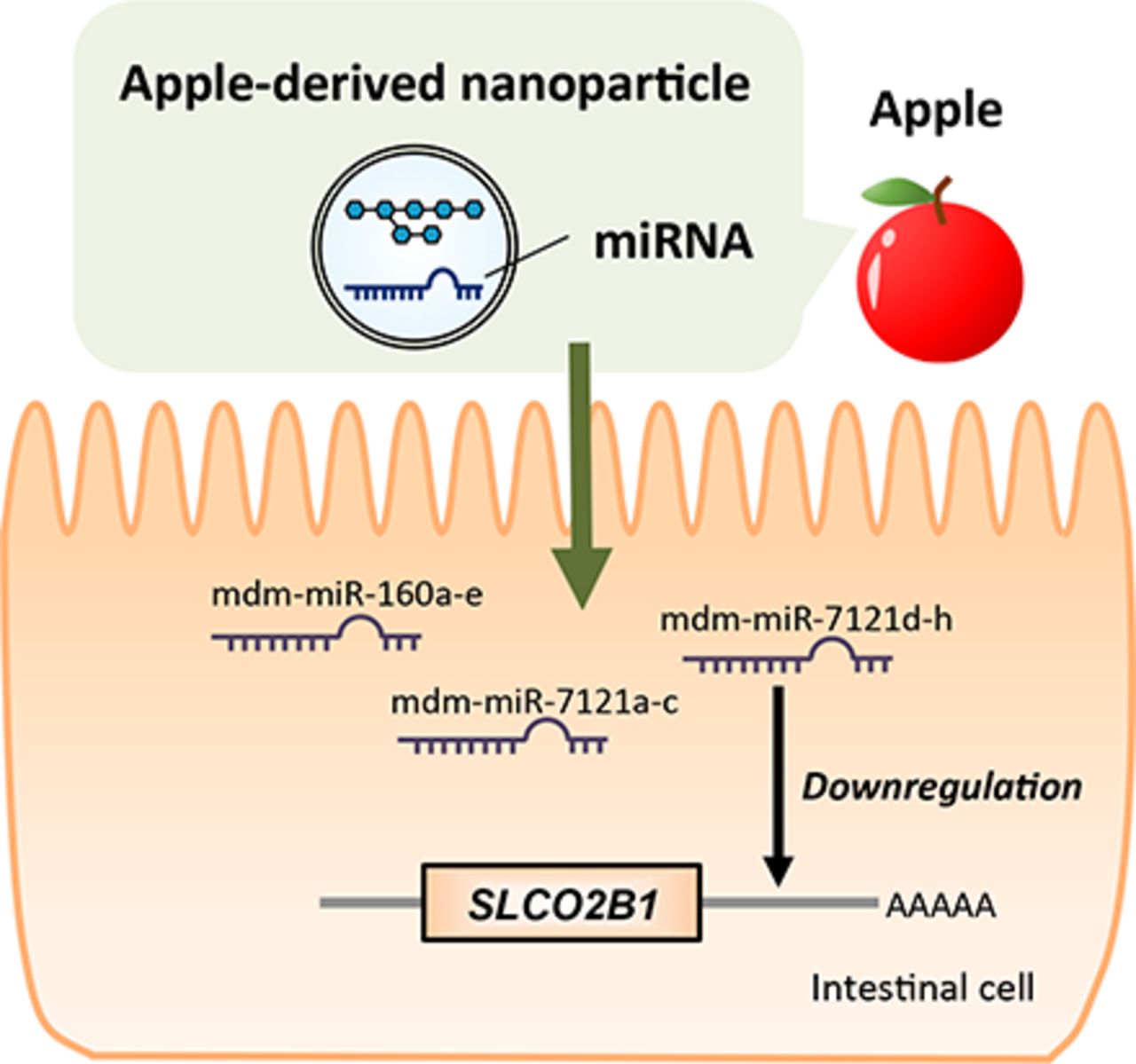
MicroRNAs in Apple-Derived Nanoparticles Modulate Intestinal Expression of Organic Anion–Transporting Peptide 2B1/SLCO2B1 in Caco-2 Cells | Drug Metabolism & Disposition

Cellular microRNA detection with miRacles: microRNA- activated conditional looping of engineered switches | Science Advances
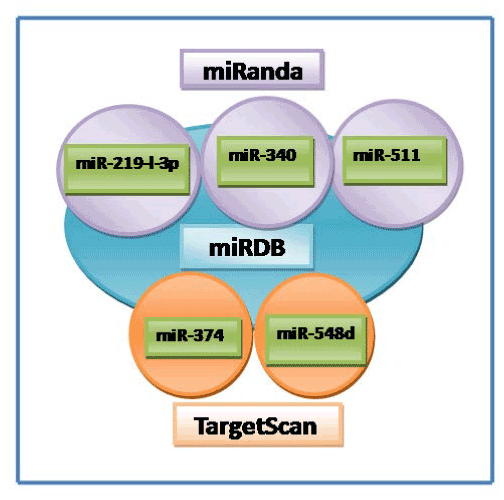
Computational prediction of novel human miRNAs/miRNA target sites in correlation with SNP (rs678653) at 3'UTR of cyclin D1 gene
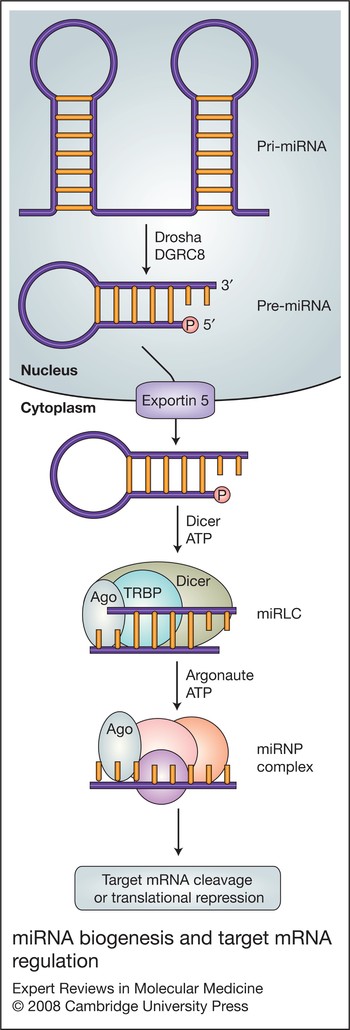
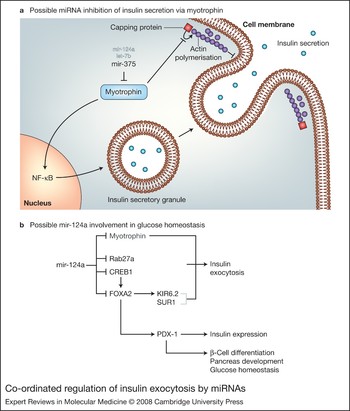
Molecular medicine of microRNAs: structure, function and implications for diabetes | Expert Reviews in Molecular Medicine | Cambridge Core
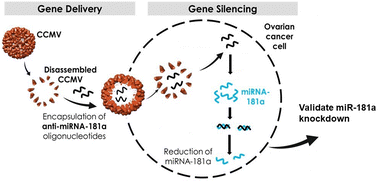
microRNA-181a silencing by antisense oligonucleotides delivered by virus-like particles - Journal of Materials Chemistry B (RSC Publishing)
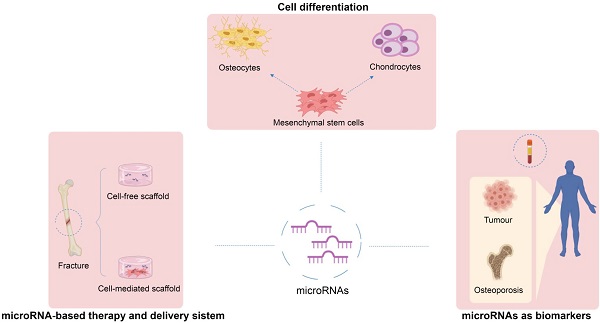
The role of microRNAs in the osteogenic and chondrogenic differentiation of mesenchymal stem cells and bone pathologies

Identification of a Plasma MicroRNA Profile Associated With Venous Thrombosis | Arteriosclerosis, Thrombosis, and Vascular Biology

High-Throughput Analysis Reveals miRNA Upregulating α-2,6-Sialic Acid through Direct miRNA–mRNA Interactions | ACS Central Science

MicroRNA dysregulation and therapeutic opportunities in esophageal diseases | American Journal of Physiology-Gastrointestinal and Liver Physiology

scanMiR: a biochemically-based toolkit for versatile and efficient microRNA target prediction | bioRxiv